- Blog
- Youtube black eyed peas i gotta feeling lyrics
- Universal audio ox
- Harry potter deathly hallows part 2 soundtrack
- Wrong turn 1 full movie watch online free
- Spider man edge of time pc release date
- Mpo sound nes emulators
- Find ms office professional plus 2013 product key
- Contra nes 30 lives free download
- Five nights at warios 2 all jumpscares
- Pes pc graphics
- Breath of wild rating
- What is hamlet about
- Fishdom h2o hidden odyssey android
- Download extra file for ben 10 protector of earth
- Treasure island beach resort
- Slic toolkit 3-3

Although the resolution of flow maps is high if we compute the MBF for each voxel and a high resolution MBF map could provide useful information such as transmural perfusion gradient, the computation time is prohibitively long and single-voxel resolution is not necessary for diagnosis. Then, for each slice, we need 15,000 to 30,000 MBF computations. In typical dynamic CT-MPI data with FOV=150mm × 150mm and 512 × 512 voxels, the area of myocardium region is 15,000 to 30,000 voxels per slice. It is time consuming to calculate MBF for each voxel on each slice for the JW model where 5 parameters need to be optimized for each voxel.
Although model-based MBF quantification may be most preferred, computation time can be a limiting factor. Previous work in CT-MPI 2 has shown that model-based deconvolution methods are more accurate and precise than model-independent methods such as singular value decomposition. MBF can be quantified from dynamic contrast-enhanced scans data with different mathematical techniques 1. Myocardial blood flow (MBF) is one of the most useful functional measurements.

SLICR even gave excellent results (103 ± 23 ml/min/100g) at 50 mAs, corresponding to 25% nominal dose.ĬT myocardial perfusion imaging (CT-MPI) can provide physiologic information for noninvasive diagnosis of cardiovascular disease. For example, at simulated MBF=100 mL/min/100g and 100 mAs exposure (50% of nominal dose) in the digital simulator, MBF estimates were 101 ± 12 mL/min/100g, 160 ± 54 mL/min/100g, and 122 ± 99 mL/min/100g for SLICR, SVD, and Johnson-Wilson, respectively. SLICR showed much better precision and accuracy than the other methods. SLICR was ~50 times faster than voxel-wise RPM and other model-based methods while retaining sufficient resolution to show all clinically interesting features (e.g., a flow deficit in the endocardial wall). We validate SLICR in a porcine model with and without partial occlusion of the LAD coronary artery with known pressure-wire fractional flow reserve.

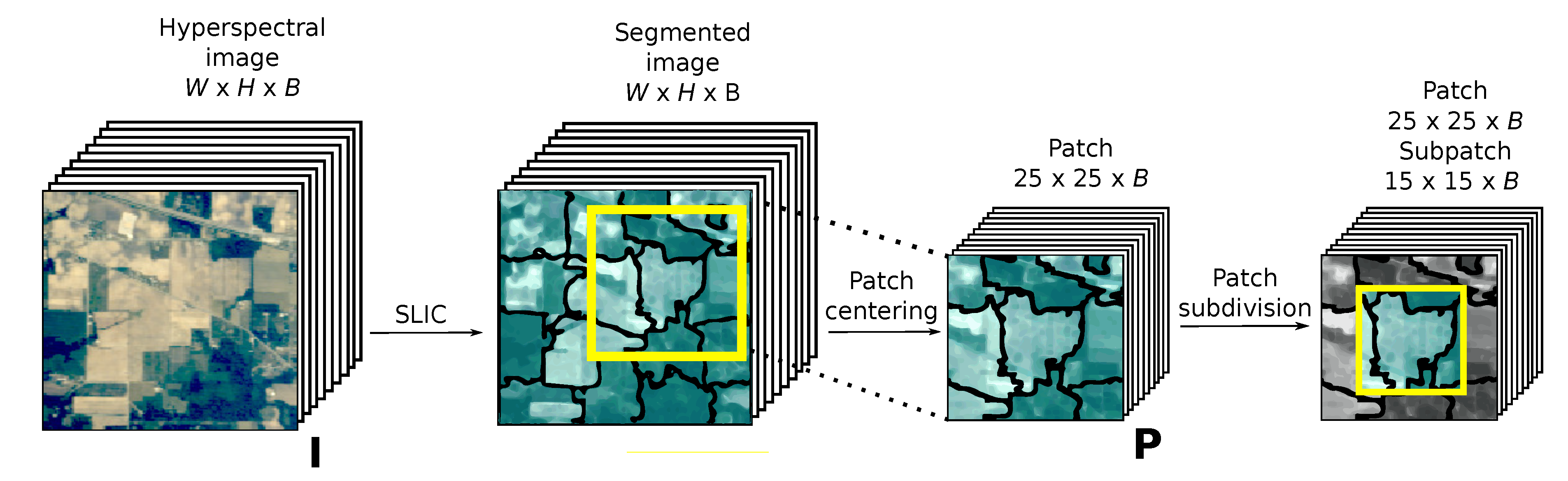
We used image data from a digital CT-MPI phantom to evaluate robustness of processing methods to noise at reduced x-ray dose. We compared SLICR against voxel-wise SVD and voxel-wise model-based deconvolution methods (RPM, single-compartment and Johnson-Wilson). We propose a new processing method, SLICR, which segments the myocardium into super-voxels using a modified simple linear iterative clustering (SLIC) algorithm and quantifies MBF via a robust physiologic model (RPM). However, iterative optimization is computationally expensive and models are sensitive to image noise, thus limiting the utility of low x-ray dose acquisitions. Previous work has shown that model-based deconvolution methods are more accurate and precise than model-independent methods such as singular value decomposition and max-upslope. There are several computational methods for estimating myocardial blood flow (MBF) using CT myocardial perfusion imaging (CT-MPI).
- Blog
- Youtube black eyed peas i gotta feeling lyrics
- Universal audio ox
- Harry potter deathly hallows part 2 soundtrack
- Wrong turn 1 full movie watch online free
- Spider man edge of time pc release date
- Mpo sound nes emulators
- Find ms office professional plus 2013 product key
- Contra nes 30 lives free download
- Five nights at warios 2 all jumpscares
- Pes pc graphics
- Breath of wild rating
- What is hamlet about
- Fishdom h2o hidden odyssey android
- Download extra file for ben 10 protector of earth
- Treasure island beach resort
- Slic toolkit 3-3
